Research
My research vision is summarized as CryptoCoLab, integrating the study of cryptogams with comparative genomics, community, collections, conservation, and collaborative science. I currently work with the themes of “evolution and ecology of urban lichens”, “hidden biodiversity discovery”, and “spatial phylogenetics and landscape genomics”.
Ongoing Research
Theme: Evolution and Ecology of Urban Lichens and Algae
Main Questions: What is the population genomic structure of urban lichens? Have lichens recolonized cities from nonurban areas or urban refugia? Are lichens evolving to survive in urban areas? What is happening with the symbiotic algae?
Project group: Jason Munshi-South (Drexel U.), Jeremy Howland (CUNY), James Lendemer (NYS Museum)
Published articles:
- Evankow et al. 2025, Urban lichens as an emerging model for urban evolution
Theme: Hidden Biodiversity Discovery
Main Questions: What undiscovered fungal and algal diversity exist in collections? What undiscovered diversity exists in understudied regions, especially the global south? Should we use single marker barcodes or genome-wide markers to study and describe biodiversity?
Project group: Research groups spanning Norway (University of Oslo, Norwegian University of Science and Technology), Sweden (Swedish Museum of Natural History), USA (Brigham Young University, U.C. Berkeley) Pakistan (University of Punjab), China (Kunming Institute of Botany), and Germany (Senckenberg Research Institute)
Published articles:
- Evankow et al. 2025, Psora mediterranea (Lecanorales, Psoraceae), a new lichen species from Europe, including a new concept for P. himalayana and a revised key to the European species
- Kirkhus et al. 2025, Diversity of tremellalean Pertusaria-associated fungi in Norway and the role of secondary metabolites in host specificity
- Brinker/Evankow Timdal 2022, Rhizoplaca ouimetensis sp. nov. (Lecanoraceae) from Ontario, the first sorediate species in the genus
Theme: Spatial Phylogenetics and Landscape Genomics
Main Questions: Where are regions with high or low levels of phylogenetic diversity? How are species distributions impacted by landscapes and other factors? Are regions of high phylogenetic diversity in conserved areas?
Project group: University of Oslo, U.C. Berkeley, and Drexel University
Preprint: Evankow et al. 2025. Revision of the deceiving lichen genus Psora in North America with descriptions of new species and spatial phylogenetic map of the continent. In: Delimiting Diversity of Lichenized Lecanorales. Faculty of Mathematics and Natural Sciences, University of Oslo, Oslo, Norway ISSN 1501-7710
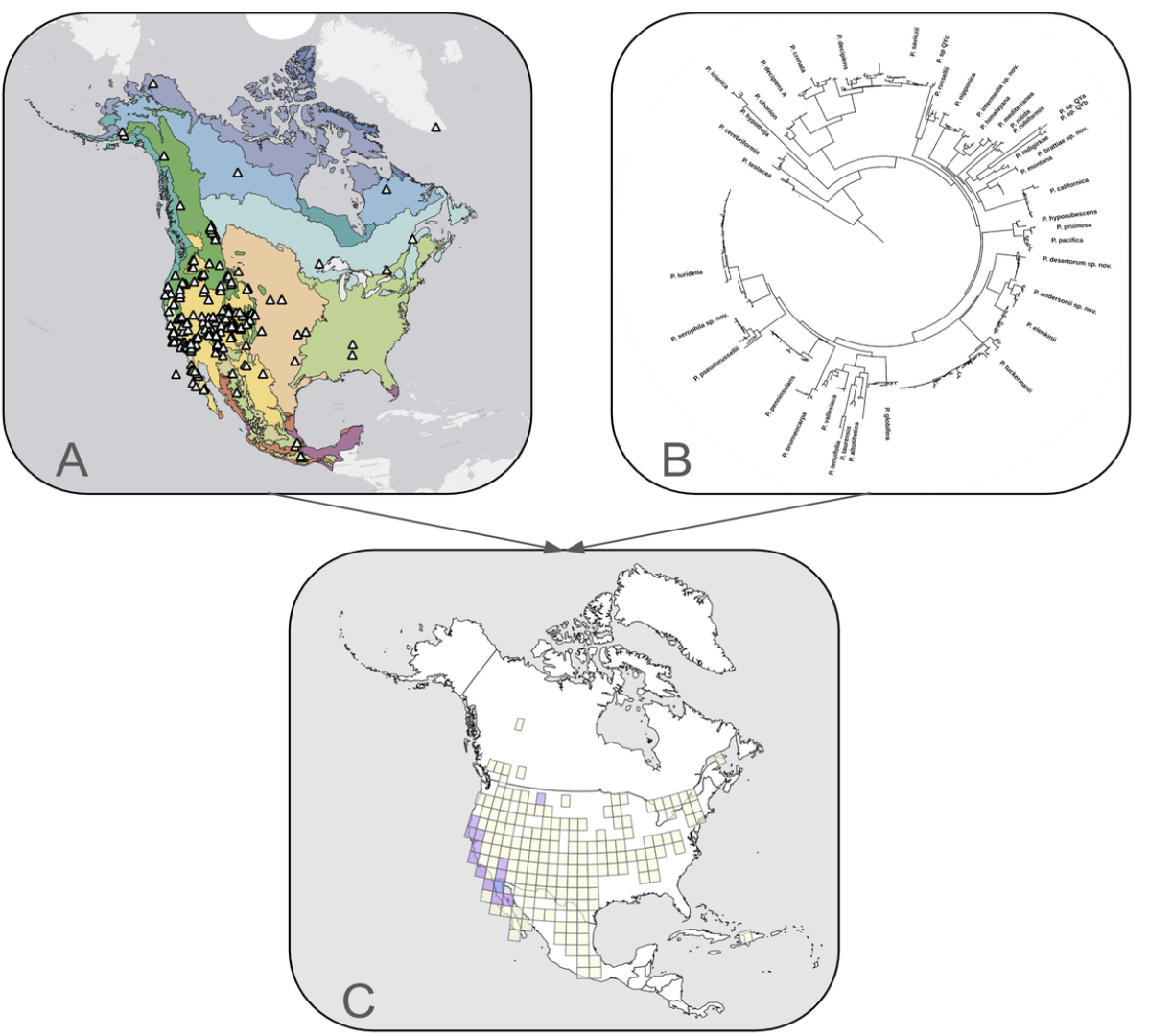 Figure Spatial Phylogenetics combines A geographical data (e.g. map of sequenced Psora specimens over North American Ecoregions) with B phylogenetic branch lengths (e.g. the phylogeny of Psora) to identify regions with elevated or low phylogenetic diversity and C Regions with mixed (purple) and paleo (blue) endemism based on CANAPE.
Figure Spatial Phylogenetics combines A geographical data (e.g. map of sequenced Psora specimens over North American Ecoregions) with B phylogenetic branch lengths (e.g. the phylogeny of Psora) to identify regions with elevated or low phylogenetic diversity and C Regions with mixed (purple) and paleo (blue) endemism based on CANAPE.
